Document Type : Original Article
Authors
1 College of Education for Pure Sciences, University of Kerbala, Iraq
2 Department of Chemistry, College of Science for Women, University of Baghdad, Iraq
Abstract
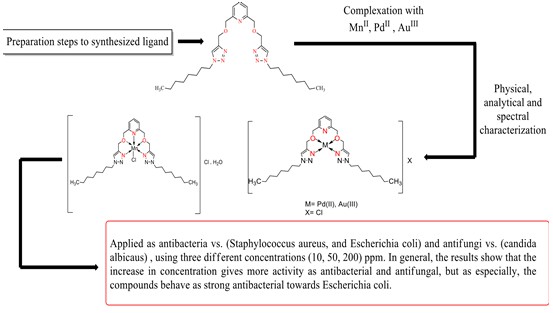
In this work, 2,6-bis(((1-decyl-1H-1,2,3-triazole-4-yl)methoxy)methyl)pyridine was prepared by click reactions between 2,6-bis((prop-2-yn-1-yloxy) methyl) pyridine and 1-azidodecane catalyzed by cuprous ion, then used as ligand in complexation reaction with some metal ions [Cu(II), Pt(IV) , and Au(III)]. The prepared ligand chelates N2O2 with copper (II) and Gold (III), as well as N3O2 with Platinum (IV). Characterized by [Flame-AAS, C.H.N. analysis, 1H-13C-NMR, Mass Spectroscopy, UV/Vis, FT-IR, conductivity, and magnetic susceptibility] were used to describe these compounds in the solid-state. The L-Cu and L-Au complexes have square planer geometry, while the L-Pt complex has octahedral geometry. The biological behaviors as antibacterial (against Staphylococcus aureus, and Escherichia coli) and antifungal (against candida albicaus) of the prepared ligand with its complexes were tested in different concentrations (10, 50, 200) ppm after incubation at 37ºC for 24h.The results showed that the performance of the prepared compounds was better in resisting and inhibiting the growth of tested bacteria and fungi at high concentrations and that the gold complex was more synergetic effective than others.
Graphical Abstract
Keywords
Main Subjects
Introduction
Triazoles are aromatic heterocycles with five members and three nitrogen atoms. These atoms can be arranged in any order, to obtain 1,2,3- or 1,2,4- isomers of triazoles. Comparing the two isomers, 1,2,3-triazole has a highly distinguished characteristic. This difference may be an illustraten illustration of this difference. This difference may be found in specialized applications of N2 substituted triazoles, such as its efficacy in cross-linking in coordination polymers to give it their mechanical properties [1]. The differences in basicity between the N1 and N2 isomers may explain their divergent behavior in biological systems, opening new avenues for pharmaceutical study. Dual orexin receptor antagonists [2], 2,3-oxidosqualene cyclases inhibitors [3], -glycosidase, and serine hydrolase are all physiologically active molecules that include the N2-1,2,3-triazole core [4]. The metal complexes with ligands (have heterocyclic moieties that include active heteroatoms such as oxygen, nitrogen, and sulfur) have recently gotten a lot of interest because of their improved bactericidal, fungicidal herbicidal, and insecticidal actions, as well as their potential as medicines [5]. This work aimed to investigate the antibacterial activity of Copper, Platinum, and Gold complexes containing ligands generated from 1,2,3-Triazole.
Materials and Methods
The chemicals were purchased from Sigma-Aldrich, Merck and were not filtered before use. The elemental microanalysis was performed using the Euro EA3000. The molar conductivity (units) of metal complexes (10-3M) in DMSO at ambient temperature was calculated using the following instruments: conduct meter (WTW), susceptibility to magnetic fields at 25°C, the complexes were measured using the Balance Johnson Mattey Catalytic System division. The Orbitrap LTQ XL ion trap MS instrument was used to obtain HRMS at positive ion mode with the source of an electrospray-ionization (ESI) at the University of New South Wales, Australia. The electronic spectra of the prepared ligand with its metal complexes were obtained by using Uv-Vis. 1800 PC in a liquid state. Shimadzu spectrophotometer with wavelengths ranging from 1100 to 190 nm. NMR spectra were acquired on 600 MHz Bruker DPX spectrometers. SHIMADZU FT-IR 8400S Fouriertrans forms with KBr and CsI discs were employed to measure FTIR-spectra in the range of wavenumber = (4000 - 200) cm-1. TLC plates made of silica were utilized in addition to an aluminum backing plate (60 F254, 0.2 mm). TLC was used to monitor the reactions, and the TLC plates were developed using an alkaline potassium permanganate dip to view them.
Synthesis of the ligand 2,6-bis (((1-decyl-1H-1,2,3-triazol-4-yl) methoxy) methyl) pyridine (L):
Preparation of this ligand according to the following steps:
A- Synthesis of n-alkyl azides (1-azidodecane) [D1]:
The solution of [1-bromodecane (10 mL) in DMF (70 mL)] was added to sodium azide (9.435 g, 0.145 mmol), the resulting suspension was continuously stirred at (60-70) °C in an oil bath for (5-6) hours after this poured the mixture into (140 mL) of distilled water and used diethyl ether (3× 50 mL) to extracted the mixture of reaction then washed with saturated organic layer 1 -azidodecane as a light yellow liquid, and purified by Colom chromatography on silica gel (Rf = 0.82 in n-hexane) to form a colorless liquid.
B- Synthesis of 2,6-bis((prop-2-yn-1-yloxy) methyl) pyridine [D2]:
In a dry flask, the DMF (50 mL) was used to dissolve pyridine 2,6-dialydiethanol (2.8 g, 20 mmol), then NaOH (3.2 g, 80 mmol) was added. The reaction mixture was chilled at the temperature (-20 °C) using an ice-cold bath, and the contents were agitated in about 10 minutes, then the propargylbromide (3.79 mL, 44 mmol) was added dropwise. After this, stirred the mixture of reaction at room temperature for 24 hours, where 100 mL distilled water was used to quench the reaction mixture, and extract it by (3×50 mL) ethyl acetate. The combined organic layer was washed off with the liquid, resulting in a pale yellow oil. As a final step, eliminate all distilled water the result is under reduced pressure using Na2SO4, filter, and evaporate the solvent until reaches dryness. The filtrate of product 2,6-bis ((prop-2-yn-1-yloxy) methyl) pyridine is (2.23 g) and the purity confirmed by using Colom chromatography on silica gel (Rf = 0. 75 in n-hexane: ethyl acetate = 9:1).
C-Synthesis of 2,6-bis-(((1-decyl-1H-1,2,3-triazol-4-yl) methoxy) methyl) pyridine (L):
The prepared compound [D2] in step B (0.215 g, 1 mmol) was dissolved in (3 mL) of DMSO and dropwise added to the heterogeneous mixture consisting of [sodium ascorbate (0.039 g, 0.2 mmol) in (3 mL) of DMSO with CuSO4.5H2O (0.025 g, 0.01 mmol) in (2 mL) of DMSO], then 1-azidodecane (3 mmol) was added to the total mixture after 2 minutes of stirring. The mixture of reactants was heated until (50-60) °C with continuous stirring for (48 hrs.). Diluted the product with 30 mL of distilled water, extracted with (3×50 mL) DCM, then eliminated distilled water residues under reduced pressure using Na2SO4, filtered, and evaporated the solvent until reached dryness. Finally, to obtain a white solid product, Colom chromatographed on silica gel (Rf = 0.17, 2:1, EtOAc: n-Hexane).
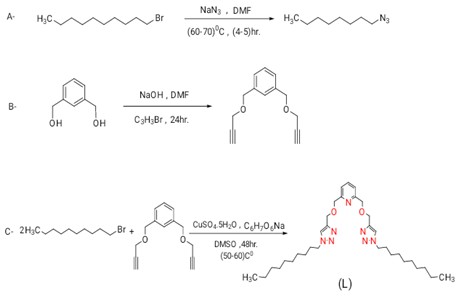
Scheme 1: Synthesis steps of the ligand (L)
Synthesis of work complexes
Each of the following metal salts (1 mmol) [CuCl2.2H2O (0.170 g), H2PtCl6 (0.486 g), and HAuCl4.H2O (0.411 g)] undergoes the reaction in an ethanoic solution. In a (1:1) molar ratio, (0.581 g, 1 mmol) of ligand was added to (10 mL) of ethanoic solution of metal salts with stirring. For 2–3 hours, the mixture was heated at reflux. During that time, a color precipitate formed, which was filtered and rinsed with ether numerous times before being dried in a desiccator. All the prepared complexes were characterized using spectroscopic, analytical, and physical measurements as shown in Table 1.
Biological activity of antibacterial and antifungal
The antibacterial activity was studied using agar diffusion technique [6]. The activity was studied versus (Escherichia coli, and Staphylococcus aureus) and candida albicaus where the method included uniformly inoculated of the surface of Moller-Hinton agar via spreading 100μl of (1×109 cells/mL) of bacterial suspension in a petri dish (90 mm diameter) for 24 h., used Macfarland solution as standard to adjust the turbidity of the bacterial suspension. Then submerge 0.1 mL of each concentration (10, 50, and 200) ppm of (prepared ligand and its complexes [L-Cu (II), L-Pt (IV), and L- Au (III)] that dissolved in dimethyl sulfoxide) into the wells (5mm diameter), and leave the plates for a half-hour after this, the plates were incubated for 24 h. at 37°C, and by measuring the inhibitory zone (mm), their ability to be antibacterial can be determined.
Results and Discussion
The isolation ligand and its prepared complexes were diagnosed based on the spectroscopic data and physio-chemical methods, that summarized in (Table 1), to suggest the general formula of the complexes as [L-Cu (II)], [L-Pt (IV)] and [L-Au (III)].

FT-IR spectra
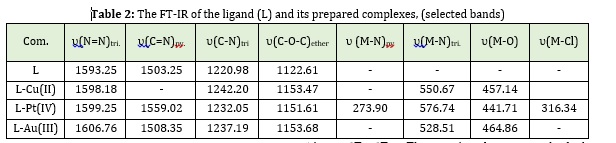
The infrared spectroscopic study has been done in the solid-state for each ligand and its complexes by employing cesium iodide. FT-IR data gave excellent information on the ligand's ability to form complexes with metal ions used in this work, and by comparison with previous literature, can specify the frequencies of ligand (as free) and its complexes (L+M) [7]. The compound 1-azidodecane was synthesized by SN2 reaction of Bromodecane with sodium azide in DMF for (4h). FT-IR spectra of compound 1-azidodecane [D1], which display the highly characteristic azide group absorption at 2094.76 cm-1, are excellent evidence for the formation of compounds 1-azidodecane [8]. The spectra of the derivative [D2] indicates the success of the reaction, where we note that the broad-band at (3354.79) cm-1, was disappeared while appearing other bands at (3308.03) cm-1 and (2121.77) have a sharp shape where it belongs to (C–H and C≡C)Terminal alkyne respectively [9]. From (Figure 1) that represent the FT-IR spectrum of the ligand (L), we can observe the (-N3 and C≡C) groups bands disappear at the region υ = (2094.76, 2117.91) cm-1 and weak bands will appear at υ = (3091.99, 1699.34) cm-1 which indicated that the aromatic triazole ring was formed and this confirms the success of cycloaddition reaction [10]. Also Figure 1 demonstrates another bands which attributed to the frequencies of these groups: υ (C-H)Pyridine = 3091.99 cm-1, υ(C=N)Pyridine = 1503.25 cm-1, υ (C-H)Triazole = 3138.29 cm-1, υ (N=N)Triazole = 1593.28 cm-1, υ(C-N)Triazole = 1220.98 cm-1, υ(C-O-C)Ether = 1122.61 cm-1 [11].
In the complexes, the frequency of this moiety υ (N=N) suffering shifted towards a higher frequency about (4-13) cm-1 and the frequency υ(C-N)Triazole, υ(C=N)Pyridine and υ(C-O-C)Ether also suffering to shifted towards a higher frequency about (12-22) cm-1, (5-56) cm-1 and (29-33)cm-1 respectively [12]. The appearance of new bundles for υ (Pt-N)Pyridine which appearance at (273.90) cm-1 in the complex of (L-Pt), also υ(M-N)Triazole which appearance at (528-576) cm-1 and, for υ(M-O) which appearance at (441-464) cm-1 in the complexes. While υ(Pt-Cl) which appearance at (316.34) cm-1[13], other bands are shown in Table 2.
Therefore, from these observations, we can have proposed that the complexes exhibit four and six coordination geometry in the solid-state. The (Figures 1-4) showed the FTIR spectra for all prepared compounds.

Electronic spectral and magnetic moment studies and Conductivity measurements
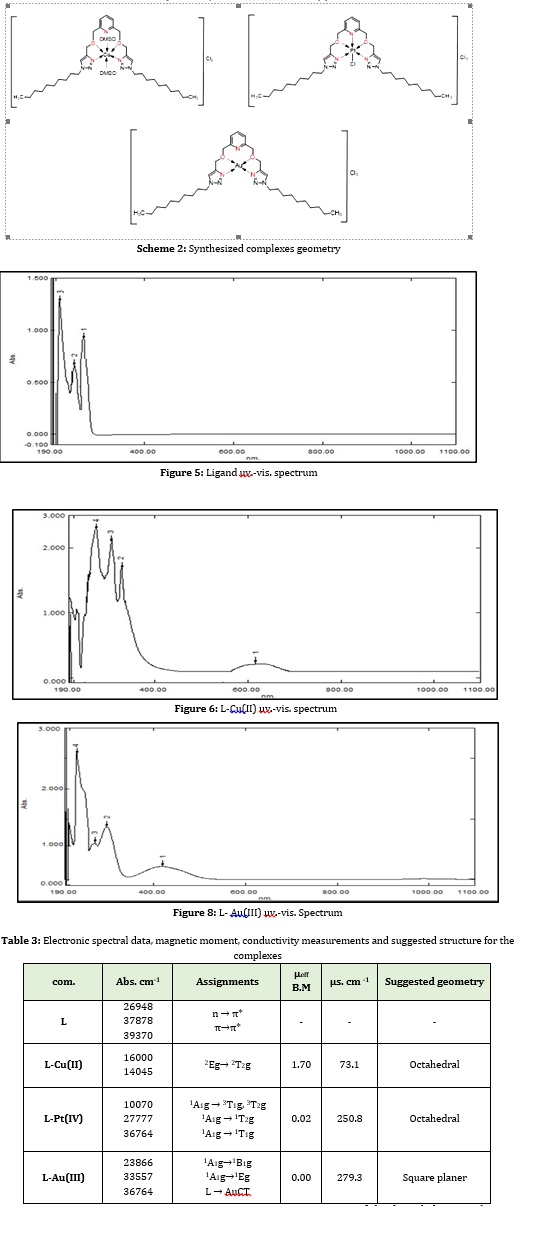
Figure 5 depicts three bands in the ultraviolet region of the linker's UV-visible spectrum (L). On the Pi system, the first band developed at 39370 cm-1 due to an internal ligand transition to (π→π*). The second absorption band, at 37878 cm-1, was raised in the same way as the transition (π→π*), although it belonged to a separate set. The electronic transition site (n → π*) on the oxygen and nitrogen atoms was connected to the third absorption band, which emerged at 26948 cm-1 [14].
[L-Cu(II)] complex:
The green complex of Cu(II) ion has a UV-VIS spectrum. During the dissolution of the solid component, the color of this complex in DMSO changed from green to brown. This shows how the geometry changed from a square planer to an octahedral structure. Figure 6 reveals the produced compound's electronic spectrum in solution, which shows a broad band from 16000 to 14045 cm-1 that corresponds to the 2Eg → 2T2g transition [15]. This compound has a magnetic value of 1.70 B.M., indicating a high spin state agreeing with the octahedral geometrical shape [16].
[L-Pt(IV)] complex
The spectrum of UV.-VIS. of prepared yellow Pt(IV) complex in Figure (7), which showed absorption bands (27777, and 36764) cm-1, which back to the transitions 1A1g → 1T2g, 1A1g → 1T1g. The band that appeared in 10070 cm-1 can be referred to as the spin forbidden transitions 1A1g → 3T1g,3T2g. These pointed to an octahedral geometry, which was confirmed by reports. The solid complex's magnetic moment was discovered to be (0.02 B.M.). This result referred to t2g6eg0 configuration spin pair octahedral stereochemistry [17]. The conductance behavior shows that this compound has an electrolyte. Furthermore, data processing and spectroscopy techniques revealed that octahedral geometry was suggested for this compound.
[L-Au(III)] complex
The charge transfers bands that dominate the ligand field transition have been used to diagnose the Au(III) ion spectrum. This means that charge transfer bands will appear at a larger wavelength while the ligand field transition will occur at a shorter wavelength. From the spectrum of this complex that shown in Figure 8, we can notice the presence of two transitions bands at 23866, 33557cm-1 which refers to 1A1g→1B1g, 1A1g→1Eg respectively [18]. Also, the spectrum indicates that charge transfer has occurred that appears as a band at 36764 cm-1. By measuring the magnetic moment of synthesized complexes, the value of μeff of this compound was equal to zero. The conductivity of this compound in DMSO at ambient temperature indicates that it is ionic. A square planar structure can be predicted for this compound based on data analysis and spectroscopy techniques. All above data of the electronic transitions and more were collected in Table 3.
1H-NMR and 13C-NMR spectra for ligand:
A) 1H-NMR spectrum
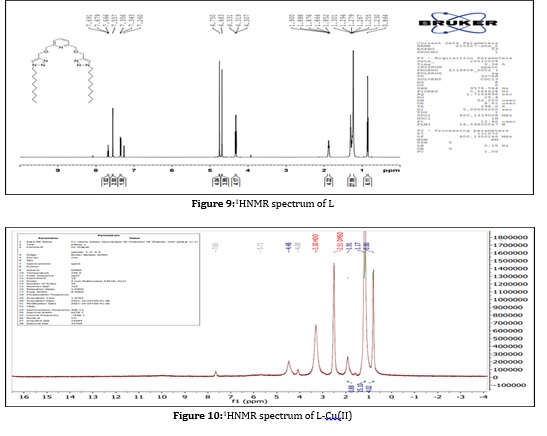
1HNMR spectrum of this ligand shown in (Figure 9) was recorded in d6-DMSO solvent and showed the following characteristic chemical shifts. Showed peaks at the range δ (1.900-0.864) ppm (d,2H, H2-H9 and H25-H32) which due to the protons of (CH2) aliphatic series groups and multiple peaks appear at the range δ (7.691-7.260) ppm (t,H, H16-H18) and (s,H11,H23) return to the (CH) groups in pyridine and triazole ring protons, and peak which appeared at the range δ(4.750-4.683) ppm (d,2H,H10,H24,H14 and H20) CH2. The first group in the aliphatic chain linked to the triazole ring and CH2 group linked the pyridine ring. Also shows the signal as peaks in δ (4.331-4.307) ppm (s, 2H, H13 and H21) [19]. The comparison of 1HNMR spectra for Cu(II) complex formed with the ligand spectrum position of proton confirms the coordination, a minimal shifting in the proton position of the ligand with these metal complexes. The DMSO exhibits singlet peaks at δ (2.48, 2.51) ppm, all shown in (Figure 10).
B) 13C-NMR spectrum
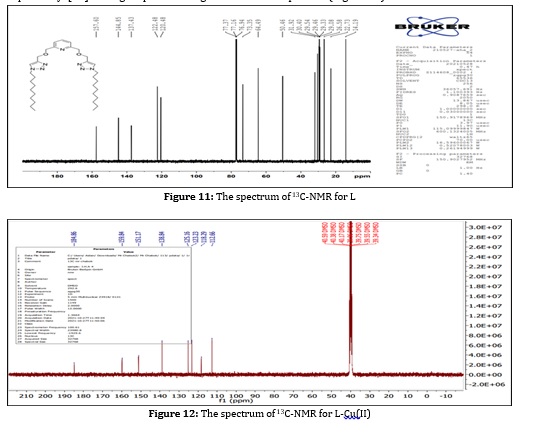
In the 13C-NMR spectrum of the ligand (L) shown in (Figure 11), shows the following characteristic chemical shift DMSO-d6 (ppm). Showed multi peaks at the range δ (31.92 - 22.73) ppm (s, C, C2-C9 and C25-C32), returning to the carbon in aliphatic series groups. While CH3 terminal group showed single peak at the position δ (14.19) ppm (s, C, C1-C29). However, the CH2. The first group in the aliphatic chain linked to the triazole ring showed one peak at the position δ (50.46) ppm (s, C, C10, C24). In contrast, CH2 group connected to the triazole ring on the other side showed one peak at the position δ (64.49) ppm (s, C, C13, C21). As for the CH2 Associated with the pyridine ring showed a peak at the range δ (77.37-73.34) ppm (s, C, C14, C20), as well CH groups of pyridine and triazole ring showed peak at δ (120.48, 122.48) ppm (s, C, C16, C18, C11, C23) and the showed peak at δ (137.43, 144.85, 157.60) (s, C, C17, C12, C22, C15, C19) respectively. The comparison of 13CNMR spectra of Cu(II) complex formed with the ligand spectrum. There is shifting in the peak of CH group of pyridine and triazole ring showed a peak at the range δ (125.16, 112.66) ppm (s, C, C16, C18, C11, C23) and the showed peak at δ (138.84, 151.17, 159.84) (s, C, C17, C12, C22, C15, C19) respectively [20]. The groups in the ligand with these metal complexes confirm the coordination compounds (Figure 12).
Mass spectra for ligand (L)
Various triazole derivatives containing different substituents have been suggested and investigated. From studying mass spectra of ligand (L) (Figure 13) show the formation of the organic ligand is confirmed by the appearance of a base peak at m\z+=604.43072 g\mol-1 corresponding to the formula [L + Na] + in HRMS which agreed well with the experimental ligand formula, (C33H55N7O2+Na) [21].

Mass spectra L-Cu(II) complex
The proposed formula corresponds to the molecular weight, (Figure 14) show the fragmentation obtained for the complex [22]. The complex formation is confirmed by the appearance of a base peak at m\z+=716.4 g\mol-1 corresponding to the formula C33H55Cl2N7O2Cu which agreed well with the experimental L-Cu(II) formula.

Antimicrobial activities
The prepared ligand and its metal complexes were screened in vitro to evaluate their action against Staphylococcus aureus as a gram-positive and Escherichia coli as a gram-negative species and antifungal activity against the antifungal activity against the antifungal activity against micro-organism candida albicaus, at three different concentrations.
The results show that most of the complexes are more toxic than the free ligand toward these bacteria and fungi at higher concentrations.
The activity of synthesized metal complexes may be due to the effect of metal ions on the normal cell membrane [23,24]. Metal chelates bear polar and non-polar properties together, making them suitable for cell permeation. And also these high activities may be referred to the Tweedy's chelation theory [25]. In addition to supposing these reasons may be due to the synergistic effect between the metal ion and organic molecule, which explain the considered hard metal ions making their complexes less lipophilic, which relatively retard their permeation through the lipid part of the cell membrance. On the other hand, the salt metal ions render their complexes to be more lipophilic, facilitate the penetration through the cell member wall, and affect the environment of the cells.
The results show complexes' import significant effect on tested gram-negative bacteria and fungal. These results may be due to multi-factors: (the nature of the metal ion, the nature, and chelate effect of the organic molecules, the nature of atoms that coordinate with metals, and the orientation of the ligand around the metal ion, nature of the metals and the oxidation state of them and the geometrical structure of these complexes.

Figure 15: Summary of antimicrobial activity of studying compounds

Conclusion
From the results obtained, it follows that new 2,6-bis (((1-decyl-1H-1,2,3-triazole-4-yl) methoxy) methyl) pyridine with its complexes were synthesized and the structure of these compounds were confirmed with spectroscopic studies, elemental analysis, molar conductivity and magnetic moment at room temperature. We suggested that Cu(II) and Au(III) complexes have four coordination number with square planer geometries, while the Pt(IV) complex has six coordination with octahedral geometry in the solid state. These compounds were evaluated for their anti-bacterial (Staphylococcus aureus, Escherichia coli) and anti-fungal (candida albicaus), and showed the synergetic effect for these complexes as high biological activity than the free ligand.
Funding
Not applicable.
Authors' contributions
All authors had equal role in study design, work, statistical analysis and manuscript writing.
Consent for publications
All authors approved the final manuscript for publication.
Conflict of Interest
The authors have no conflict of interest to declare.
ORCID
Jihan Hameed Abdulameer:
https://www.orcid.org/0000-0003-2850-8458
HOW TO CITE THIS ARTICLE
Jihan Hameed Abdulameer, Mahasin F. Alias, Synthesis, Characterization and Antimicrobial Activity of Cu (II), Pt (IV) and Au (III) Complexes with 2,6-bis (((1-decyl-1H-1,2,3-triazole-4-yl) methoxy) methyl) pyridine. Chem. Methodol., 2022, 6(3) 184-196
DOI: 10.22034/CHEMM.2022.318629.1405
- Bhagat U.K., Peddinti R.K., Org. Chem., 2018, 83:793 [Crossref], [Google scholar], [Publisher]
- Bozorova K., Zhaoa J., Aisa H.A., Med. Chem., 2019, 27:3511 [Crossref], [Google scholar], [Publisher]
- Martín J, Alés RM, Asuero AG. An overview on ligands of therapeutically interest. Pharm. Pharmacol., 2018, 6:198 [Google scholar]
- Lopes A.B., Wagner P., Kummerle A.E., Bihel F., Bourguignon J.J., Schmitt M., Miranda L.S.M., Select, 2017, 2:6544 [Crossref], [Google scholar], [Publisher]
- Gavlik K.D., Lesogorova S.G., Sukhorukova E.S., Subbotina J.O., Slepukhin P.A., Benassi E., Belskaya N.P., J. Org. Chem., 2016, 2016:2700 [Crossref], [Google scholar], [Publisher]
- Zhang S., Xu Z., Gao C., Ren Q.C., Chang L., LV Z.S., Feng L.S., J. Med. Chem., 2017, 138:501 [Crossref], [Google scholar], [Publisher]
- Zhu C., Zeng H., Chen F., Liu C., Zhu R., Wu W., Jiang H., Chem. Front., 2018, 5:571 [Crossref], [Google scholar], [Publisher]
- Meinel R.S., das Chagas Almeida A., Stroppa P.H.F., Glanzmann N., Coimbra E.S., da Silva A.D., Biol. Interact., 2020, 315:108850 [Crossref], [Google scholar], [Publisher]
- García-Monroy R., González-Calderón D., Ramírez-Villalva A., Mastachi-Loza S., Aguirre-de Paz J.G., Fuentes-Benítes A., González-Romero C., Mex. Chem. Soc., 2021, 65:202 [Crossref], [Google scholar], [Publisher]
- Motika S.E., Shi X., ARKIVOC, 2018, 2018:280 [Crossref], [Google scholar], [Publisher]
- Kaur S., Kaur P., Indo Glob. J. Pharm. Sci., 2019, 09:146 [Google scholar]
- Joy M.N., Bodke Y.D., Telkar S., Bakulev V.A., Mex. Chem. Soc., 2020, 64:46 [Crossref], [Google scholar], [Publisher]
- Al-Radadi N.S., Zayed E.M., Mohamed G.G., Abd El Salam H.A., Chem., 2020, 2020 [Crossref], [Google scholar], [Publisher]
- Mohamad H.A., AL-Kattan W.T., AL-Daly Z.M., Chemistry, 2020, 36:903 [Google scholar], [Publisher]
- Karczmarzyk Z., Marta Swatko-Ossor M., Wysocki W., Molecules, 2020, 25:1 [Crossref], [Google scholar], [Publisher]
- Carver P.L., Metal Ions Life Sci., 2019, 19:1 [Crossref], [Google scholar], [Publisher]
- Al-Hamdani A.A.S., Hamoodah R.G., Baghdad Sci. J., 2016, 13:770 [Google scholar], [Publisher]
- Shaker S.A., Mohammed H.A, Al-Hamdani A.A.S., J. Basic Appl. Sci., 2010, 4: 5178 [Google scholar]
- Noor M.W., Al-Hamdani A.A.S., Al-Zoubi W., Phys. Org. Chem., 2020, 33:1 [Crossref], [Google scholar], [Publisher]
- Al Zoubi W., Al‐Hamdani A.A.S., Duraid Ahmed S., Basheer H.M., Al‐Luhaibi R.S., Dib A., Ko Y.G., Phys. Org. Chem., 2019, 32:3916 [Crossref], [Google scholar], [Publisher]
- Ahmed S.M., Salih K.M., Ahmad H.O., Jawhar Z.H., Hamad D.H., Zanco J. Med. Sci., 2019, 23:206 [Crossref], [Google scholar], [Publisher]
- Al-Hamdani A.A.S., Al-Luhaibi R.S., RJPBCS, 2017, 8:164 [Google scholar]
- Anitha C., Sumathi S., Tharmaraj P., Sheela C.D., J. Inorg. Chem., 2012 Jan 24; 2011:1 [Crossref], [Google scholar], [Publisher]
- Abouzayed F.I., Emam S.M., Abouel-Enein S.A., Mol. Struct., 2020, 1216:128314 [Crossref], [Google scholar], [Publisher]
- Kumari S.S., Asian J., 2020, 32:192 [Google scholar], [Publisher]