Molecular Docking and Dynamic Simulation with ADMET Exploration of Natural Products Atlas Compound Library to search for Potential Alpha Amylase Inhibitors for the T2DM Treatment
Pages 81-102
https://doi.org/10.48309/chemm.2025.488471.1849
Venugopal Muralidharan, Rahul Ashok Sachdeo, Laliteshwar Pratap Singh, M. Akiful Haque, Uttam Prasad Panigrahy, Abhishek Kumar Mishra, Touseef Begum, Mirza Shahed Baig
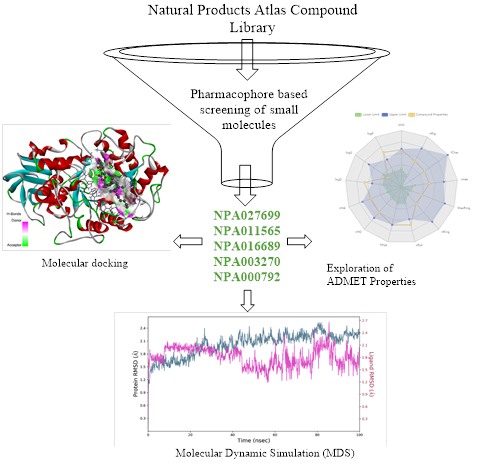
Abstract One of the major global causes of early mortality and illness is diabetes. Diabetes mellitus, a global metabolic condition, affects almost every age group in the world's population. The primary cause of diabetes is insufficient pancreatic function, which occurs when the organ fails to produce enough insulin or fails to adequately use the insulin it produces. The aim of this study is to identify a lead drug as a potent alpha-amylase inhibitor to fight T2DM. We determine the pharmacological alpha amylase target (PDB ID: 3BAX) through extensive analyses and reviews of the available literature. We used a pharmacophore query to search the Natural Products Atlas library for a potent inhibitor. The PyRx version of AutoDock Vina is utilized for the docking process. We use the Desmond software to identify the stability of complexes in physiological environments. According to our analysis, the primary substances NPA016689 and NPA011565 are potent inhibitors against alpha-amylase (3BAX). After determining the lead, we identified the NPA016689 and NPA011565 compounds as the most reactive, with binding affinities of -9.3 kcal/mol and -9.2 kcal/mol, respectively. TYR_A:62, TYR: A_151, LYS: A_200, HIS: A_201, ILE: A_235, and HIS: A_305 residues critically interacted with the NPA016689 ligand. Similarly with ligand NPA011565, the binding residues were TRP: A_59, THR: A_163, LEU_A:165, ARG: A_195, ASP_A:197_A:198, GLU: A_233, and HIS: A_299. This study concluded that NPA016689 and NPA011565 have the potential to modulate the activity of alpha-amylase enzyme, making them potential lead molecules for the design or development of additional anti-alpha-amylase compounds.