Document Type : Original Article
Authors
1 Faculty of Pharmacy, M.S. Ramaiah University of applied sciences, M.S.R.I.T. Post, Bangalore – 560054, Karnataka, India
2 PES college of pharmacy, Hanumanthnagar, Bangalore – 560050, Karnataka, India
Abstract
Candida albicans is an opportunistic fungal pathogen that causes candidiasis in human hosts. Candidiasis includes a multitude of fungal infections, including invasive fungal infections, where most patients are immunocompromised; hence, the success of treatment is determined by the efficacy of the antifungal agent. However, with the increase in resistance to the existing drugs, the availability of effective antifungal agents is becoming scarce.
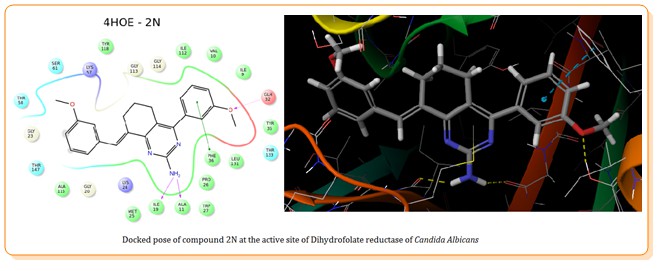
Many pyrimidine derivatives exhibit powerful antifungal activity. In this study, In silico antifungal activity was carried out on twenty novel aminopyrimidine derivatives to identify the specificity of the pyrimidine analogues for the antifungal targets using ‘Glide’. Molecular docking studies were conducted on two antifungal targets; Dihydrofolate reductase of C. albicans (PDB ID: 4HOE); N-myristoyl transferase of C. albicans (PDB ID: 1IYK); energy minimization of title compounds was carried out using LigPrep, the protein targets were optimized and minimized, a 3-dimensional grid was generated at the active site, and molecular docking was carried out at both the standard precision (SP) and extra precision (XP) modes. The docking poses were ranked according to their docking scores (GScore) and their binding energy with the enzyme (Emodel). The obtained results for the docking of the title compounds with dihydrofolate reductase of C. albicans are quite promising. Molecular docking suggest that compounds 2N and 2A are potential inhibitors of dihyfrofolate reductase and are specific in binding at the active site of the enzyme. They form pi-pi stacking interactions with PHE 36 at the active site of the protein, similar to the standard drug. However the test compounds show lower docking scores against N-myristoyl transferase of C. albicans indicating that they may not be effective against the fungal protein.
Graphical Abstract
Keywords
Main Subjects